
Review
on advances in biodegradation of phenols: kinetics, modelling and mass transfer
Ladi Reshma 1![]()
![]() ,
V. Sridevi 2
,
V. Sridevi 2![]()
![]() ,
M. N. N. Sai Rachana 1
,
M. N. N. Sai Rachana 1![]()
![]() ,
J. Akhila 1
,
J. Akhila 1![]()
![]() ,
M. Yamini 1
,
M. Yamini 1![]()
![]() ,
Katru Ramya Sugandhi 1
,
Katru Ramya Sugandhi 1![]()
![]() ,
Husam Talib Hamzah 3
,
Husam Talib Hamzah 3![]()
![]() ,
R. Sri Harsha 1
,
R. Sri Harsha 1![]()
![]()
1 M. Tech Biotechnology, Department of Chemical Engineering, AU College of Engineering (A), Andhra University, Visakhapatnam, India
2 Professor, Department of Chemical Engineering, Andhra University, Vishakhapatnam-530003, India
3 Ph.D. Scholar, Department of Chemical
Engineering, Andhra University, Vishakhapatnam-530003, India
|
|
ABSTRACT |
||
|
Harmful
pollutants like phenol and its derivatives are found in wastewater from a
wide range of industries, including oil refining, medicines, coal conversion,
chemistry, and petrochemistry. The high concentration, high toxicity, and
difficult-to-degrade characteristics of phenols in wastewater pose a serious
threat to the environment and to human health. By employing different strains
of microorganisms and biocatalysts to create biodegradation procedures of
diverse pollutants and a wide spectrum of hazardous compounds, biotechnology
has successfully addressed significant environmental challenges. Among
various phenols removal techniques, biodegradation is both economical and
environmentally friendly. During the study of microbial degradation
processes, there is a great deal of interest in the potential for
mathematical modelling to forecast microbial growth and degrade harmful or
inhibiting environmental pollutants at variable quantities. Such mathematical
models are frequently created using aromatic compounds like phenol. The
review discusses the following topics: kinetics, modelling, and mass
transfer; future scope and directions; diverse microorganisms, bioreactors,
the metabolic pathway of phenol, influencing factors, and recent advancements
in biological therapy. |
|||
|
Received 11 December 2022 Accepted 12 January 2023 Published 31 January 2023 Corresponding Author Ladi Reshma, ladi.reshma123@gmail.com DOI 10.29121/granthaalayah.v11.i1.2023.4968 Funding: This research
received no specific grant from any funding agency in the public, commercial,
or not-for-profit sectors. Copyright: © 2023 The
Author(s). This work is licensed under a Creative Commons
Attribution 4.0 International License. With the
license CC-BY, authors retain the copyright, allowing anyone to download,
reuse, re-print, modify, distribute, and/or copy their contribution. The work
must be properly attributed to its author.
|
|||
|
Keywords: Biodegradation, Phenol, Mechanism,
Immobilization, Modelling, Kinetics, Mass Transfer HIGHLIGHTS · The manufacturing of paper, coking, insecticides, plastics, and pharmaceuticals are only a few of the industrial processes that produce phenol, which has a very broad range of sources. · An intricate process involving the creation of several enzymes is required to biodegrade the aromatic molecule phenol. · When organic matter, in this case phenolic compounds, is transformed into carbon dioxide, this process is known as carbon mineralization. · To explore the kinetics of phenol biodegradation in the SBR mode and to compare the findings with the available data, the Haldane equation (Eq. This formula is used to calculate the rate at which an inhibiting substrate degrades. |
|||

1. INTRODUCTION
As industrial production increased and the demand for chemicals increased, massive amounts of phenol-tainted effluent were discharged into waterways. Phenol and its derivatives were chosen due to their extensive industrial use and threat to aquatic life Duan et al. (2018). The removal of various dangerous organic compounds from wastewater can be accomplished using three main techniques: non-destructive, oxidative destructive, and biologically destructive. Since non-destructive methods rely on the physical adsorption and removal of contaminants, they produce a lot of waste. Oxidative destructive methods include incineration, moist oxidation, and complex oxidation processes (AOPs). Although oxidative destruction may be able to eliminate contaminants, the cost of the necessary equipment, operations, and maintenance make it difficult to use. Biological activities consume a lot less energy and chemicals than other types of processes.
One of the biggest challenges to microbial pollutant degradation is the inability to retain enough biomass for the biodegradation of a target molecule. Antimicrobials can be used to treat both inhibiting compounds like phenols and microbiological illnesses in living things. If either is present, it may cause a die-off of microorganisms in the bioreactor and ineffective wastewater treatment. Although maintaining the bioaugmented culture in a bioreactor can be difficult, adding certain bacterial cultures to a system (bioaugmentation) may expedite the breakdown of antimicrobial contaminants and resistant chemicals Kuc et al. (2022). The washout and dilution of the suspended bacterial culture in a bioreactor are frequently brought on by the continuous injection of wastewater to the reactor. To get around these difficulties in biological degradation, researchers have created a variety of methods for immobilising microbial cultures as well as methods for accelerating phenol degradation through the use of kinetics, modelling, and mass transfer.
2. ORGANIC POLLUTION OF PHENOL
Today's
population must regularly deal with a frustrating issue that includes land
pollution, water contamination, other environmental problems, and air pollution
Sun et al. (2022). One of the most pressing
environmental problems on a global scale is organic pollution, which has risen
quickly in importance due to the rapid development in urbanisation and
industrialization Touliabah et al. (2022). Water that has been tainted
with phenolic pollutants is challenging to clean up since they arrive in a
variety of concentrations from various industrial processes. Phenolic compounds
are present in wastewater and are dangerous to both people and the environment
because of their high solubility in water and sluggish rate of biodegradation.
for a variety of reasons. Even at modest doses, phenols and phenolic compounds
are toxic to human health, and many of them are considered dangerous pollutants
as a result, Aminophenols, butylhydroxytoluene,
nonylphenols, and bisphenols are only a few examples of the phenolic substances
Aisami et al. (2020).
1) SOURCE
OF PHENOLS
Typically, the primary environmental sources of phenol are man-made or natural materials. It results from its processing and use in many contexts, such as, for example, wood burning and automotive exhaust Khleifat et al. (2007). Endogenous and exogenous phenolic compounds are the two types of phenolic compounds that are frequently found in nature Sun et al. (2022). Endogenous phenols are phenolic chemicals that are present in nature and have biological effects. They frequently include the leaves, stems, fruits, roots, and other plant or crop elements. The by products are exogenous phenols. Exogenous phenols and substituted phenols, mostly from vehicle exhaust and smoke from burning biomass, were present in the air. Related phenolic compounds are also created during the production and processing of industrial items. During the coking and refining processes, phenol was one of the principal ingredients obtained from coal and oil mines Wang et al. (2011), Viraj et al. (2017). The creation of oil and its by products, fibreglass, steel, furnace coke, cork, explosives, paint, the manufacture and recycling of rubber products, the textile industry, and several segments of the food and beverage industry are just a few industrial processes that produce phenols Bhatia et al. (2018).
2) LIMITS
OF PHENOL
The
EPA has established a threshold for phenol in surface water of fewer than 1 ppb
Kazemi et al. (2014). The toxicity thresholds are
typically between 9 and 25 mg/L for both aquatic life and humans. Phenol may
have immediate or long-term negative consequences on one's health. Humans who
are exposed to toxic compounds over an extended period of time may have deadly
dosages of respiratory risks, tremor, weakness, and erratic breathing. The
National Institute for Occupational Safety and Health (NIOSH REL), which stands
for "NIOSH Recommended Exposure Limit," specifies the upper limit or
exposure limit for an 8- or 10-hour timeweighted average. The acceptable
exposure limit, or OSHA PEL, is the maximum concentration of a material to
which most employees can be exposed without suffering adverse consequences. It
is a time-weighted average across a normal 8-hour workday or 40-hour workweek.
The United States Environmental Protection Agency (USEPA) recommends that lead
levels in drinking water not go over 0.05 mg/l Kazemi et al. (2014).
3) HARMFUL
EFFECTS OF PHENOL
In
addition to cancer and genetofibre striation, severe
phenol exposure produces disorders of the central nervous system, hepatic
damage, anorexia, cutaneous rash, dysphasia, gastrointestinal disturbance,
vomiting, weakness, and weightlessness González et al. (2001). According to an animal study,
oral phenol exposure results in decreased foetus weights, delayed growth, and
aberrant development in the offspring of the animals. Increased maternal
mortality and decreased maternal weight gain were also discovered González et al. (2001).
2.1. ANALYSIS OF PHENOL
Numerous techniques, including
spectrophotometry, HPLC, GC, and their combinations, have been employed to
measure phenolic chemicals from plant materials as analytical science has
advanced.
2.1.1. SPECTROPHOTOMETRY
Spectrophotometry,
which largely relies on many measurement techniques for the different
structural changes of the phenolic compounds, is a rapid and simple methodology
for figuring out how much phenolic compounds are present in plant materials.
Typically, the content of flavonoids is determined using spectrophotometry.
(2016) Pouraboli et al. Additionally, condensed
tannin concentration and total phenolic amount can both be determined using
spectrophotometry Sankhalkar and Vernekar
(2016). Spectroscopy is a frequently
utilised technique for quantifying many different types of phenolic compounds
due to its simplicity of use and inexpensive cost.
2.1.2. GAS CHROMATOGRAPHY (GC)
GC is a useful method for separating, identifying,
and measuring the numerous phenolic chemicals present in plants, including
anthocyanins, flavonoids, and tannins. Samples are heated in a heated column
that uses the evaporation temperature specific to each compound to separate it
from the solution. The column is lined with a thin layer of non-volatile liquid
that is coated with an inert substrate Vaičiulyte et al. (2016).
2.1.3. High-Pressure Liquid Chromatography (HPLC)
The
most popular method for separating and detecting phenolic chemicals is HPLC. It
is a flexible and adaptable tool with a number of benefits, including good
selectivity, sensitivity, resolution, and sample behaviour Naczk and Shahidi (2006). The basic idea behind the
method is to separate chemicals from complex mixtures based on how soluble they
are and/or how they interact with a less polar stationary phase and a more
polar mobile phase Coskun (2016). Thus, some variables,
including column types, used detectors, mobile phase, and the characteristics
of the tested substances, have an impact on HPLC analysis of phenolic
compounds.
2.1.4. HPLC–Mass Spectrometry
HPLC
and tandem MS can be used to examine phenolic substances. An innovative
analytical method with great sensitivity and selectivity is HPLC supplemented
by MS detection. With this method, unidentified chemicals in samples of natural
sources that have been partially or crudely purified can have their structural
information measured. Mocan et al. (2014). Numerous studies on the
analysis of phenolic compounds have recently concentrated on the evaluation of
techniques including various couplings between HPLC and MS.
2.1.5. HPLC–Diode Array Detector
Another
popular technique for determining the presence of phenolic compounds in plants
is HPLC combined with a diode array detector (HPLC-DAD). Da Silva Siqueira et al. (2016), Alqahtani et al. (2015).The most expensive and uncommon
of the detectors used in conjunction with HPLC to the
identification of phenolic chemicals
is MS, whereas the most practical and widespread is DAD Rejczak and Tuzimski
(2017).The entire UV/visible spectrum
of the analytes can be simultaneously scanned by the DAD detector, which can
also provide details on unique spectral characteristics for compound
identification.
3. METABOLIC PATHWAYS FOR PHENOL
An
aromatic hydrocarbon called phenol is broken down by a variety of bacteria, which
obtain all of their carbon needs from phenol. Both
aerobic and anaerobic environments can lead to the degradation of phenol.
3.1. AEROBIC BIODEGRADATION OF PHENOL
Oxygenation
starts the biodegradation of phenol under aerobic conditions. A monooxygenase
phenol hydroxylase first monohydroxylates the
aromatic ring in this process at an ortho location to the pre-existing hydroxyl
group to produce catechol. This is the primary intermediate produced when
various microbial strains break down phenol. Depending on the strain, catechol
is either oxidised by catechol 1,2-dioxygenase via the ortho-cleavage pathway,
which results in the creation of succinyl Co-A and acetyl Co-A, or by catechol
2,3-dioxygenase via the meta-pathway, which results in the formation of
pyruvate and acetaldehyde Patil et al. (2014).
3.2. Anaerobic biodegradation of phenol
The aerobic
phase of phenol decomposition is more advanced than the anaerobic process. The
aerobic phase of phenol decomposition is more advanced than the anaerobic
process. The first step in the anaerobic process is the 4-hydroxy benzoate
carboxylase-mediated carboxylation of phenol at the para position to
4-hydroxybenzoate. Carboxylation and subsequent dihydroxylation are the
mechanisms through which phenolic chemicals, such as o-cresol, catechol, and
ortho-halogenated phenol, are degraded by anaerobic bacteria González et al. (2001). Figure 1 and Figure 2 shows aerobic and anaerobic
degradation pathway for phenol.
Figure
1
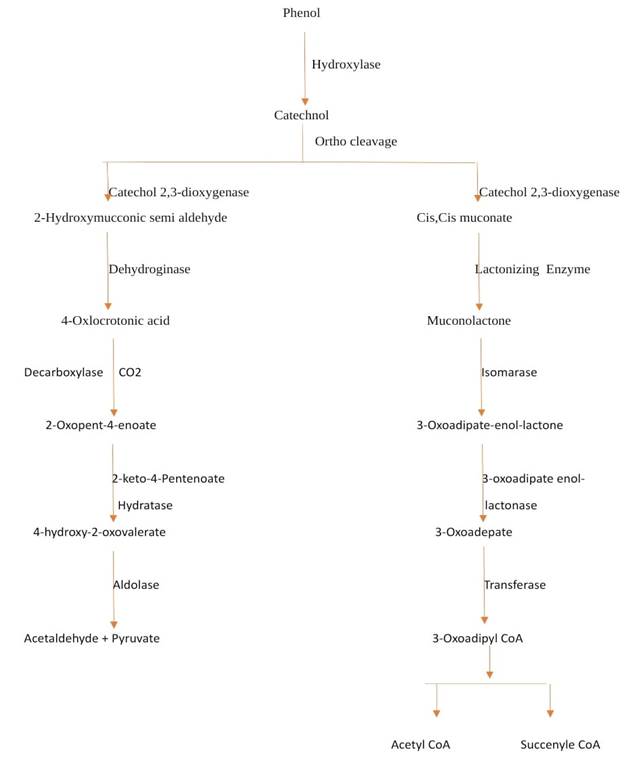
|
Figure 1 Aerobic Degradation Pathway for Phenol |
Figure 2
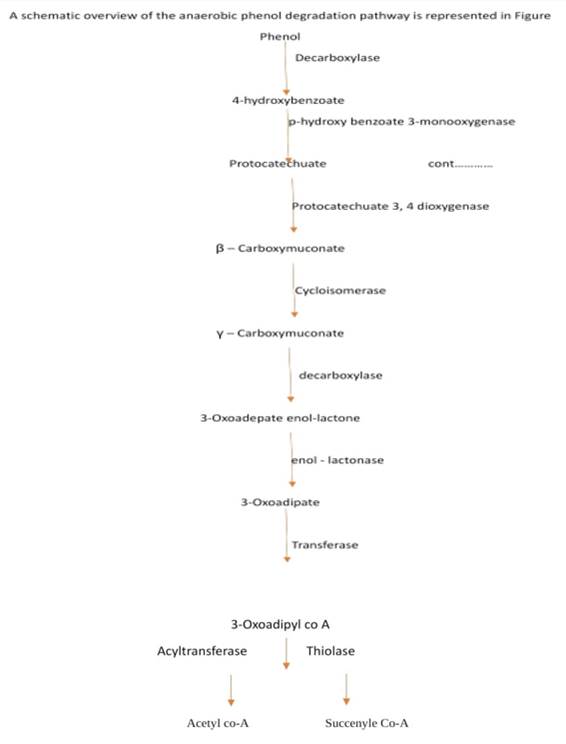
|
Figure 2 Anerobic Degradation Pathway for Phenol |
4. MECHANISM, VARIOUS MICROORGANISMS, AND REACTORS FOR PHENOL BIODEGRADATION
Phenol, an aromatic
hydrocarbon is degraded by various microorganisms, Table 1 which utilizes phenol as the sole carbon
source for the growth of the organisms. Among the various microorganisms Pseudomonas
putida is the most popular organism for the degradation of phenol as this
species uses phenol as the carbon source . Numerous
microorganisms—including both aerobic and anaerobic ones—use phenol as their
only source of carbon and energy as a result of the
substance's extensive prevalence in the environment. The presence of phenolic compounds in water
and soil has become significant problems. Common commercial wastewater
treatment methods utilize the combination of physico-chemical
and biological treatment. Both chemical and biological processes were used for
many years to treat phenolic wastewater. Activated sludge, fluidized, packed
bed and moving bed biofilm reactors were studied as biological treatment
processes. Table 2 shows various reactors used in phenol
degradation and their effect. The degradation rate depends on the state of
biomass development, feed concentration, liquid flow rate, and air flow rate.
4.1. DEGRADATION OF PHENOL THROUGH AEROBIC AND ANAEROBIC PATHWAYS
4.1.1. AEROBIC BIODEGRADATION OF PHENOL
At
the beginning of the 19th century, research on aerobic biodegradation began.
The enzyme phenol hydroxylase uses molecular oxygen to add a second hydroxyl
group in ortho-position to the one that already exists in the initial phase of
the aerobic route for the biodegradation of phenol. Pyridine nucleotide
reduction is necessary for the process (NADH2). Depending on the causative
bacterium, one of two procedures can then be used to get rid of the resultant
catechol (1, 2- dihydroxy benzene) molecule. A catechol 1, 2-dioxygenase splits
the aromatic ring between the catechol hydroxyls via the ortho- or ketoadipate
pathway (intraradiol fission) Harwood and Parales (1996). The first to provide
circumstantial evidence that strain "Vibrio 01" generated
-ketoadipate while metabolising phenol was Evans and Kilby Evans (1947).
The
resultant cis, cis muconate undergoes further
metabolism to create -ketoadipate, which is a Krebs cycle intermediate. Ring
fission takes place in the meta-pathway close to the two hydroxyl groups of
catechol (extra diol fission). Catechol 2, 3-dioxygenase is an enzyme that
changes catechol into 2-hydroxymuconic semialdehyde. This chemical is converted
into Krebs cycle intermediates by further metabolism. Acinetobacter calcoceticus, Pseudomonas species, and Candida tropicalis
use the aerobic pathway to consume phenol, whereas other eukaryotes frequently
use the ortho pathway. Numerous studies have focused on the aerobic Pseudomonas
species, and their capacity to grow on a range of aromatic substrates makes
them an appealing organism for use in wastewater treatment applications Kilby (1948).
4.1.2. Anaerobic biodegradation of phenol
The
first stage of the anaerobic phase of this process is the carboxylation of
phenol at the para position to 4 hydroxybenzoate. The 4-hydroxybenezoate
carboxylase is the enzyme in question here. It has been demonstrated that a
carboxylation reaction is a key component of the anaerobic decomposition of
many more aromatic compounds. It has been proposed that o-cresol can be denitrified
to produce 3-methyl 4-hydroxybenzoate by carboxylating
the aromatic ring in para position to the hydroxy group. Studies demonstrated
that Para coccus-like organisms and the methogenic
consortia moved through a variety of phenolic chemicals, including catechol,
ortho halogenated phenols, and o-cresol, after para Dehydroxylation,
followed by carboxylation.
Table 1
|
Table 1 Phenol Degrading Microorganisms Fan et al. (2008) |
||
|
Source |
Genus |
Species |
|
Bacteria |
Alcaligenes Anthrobacter Pseudomonas Cyanobacterium
Bacillus |
Alcaligenes
faecalis Alcaligenes
xylosoxidans Y234 Arthobacter species Arthobactercitreus Arthobacterchlrophenolicus A6 Pseudomonas
putida Pseudomonas
cepacian Pseudomonas
pictorum Pseudomonas
aeruginosa MTCC 4996 Pseudomonas
aeruginosa Pseudomonas
aeruginosa CC7CCAB919095 Cyanobacterium
synechococcus Bacillus
species strain PHN1 Bacillus
brevis Bacillus
badius |
|
Fungi |
Candida Fusarium Graphium Ochromonas Aspergillus |
Candida
Tropicalis Candida
Tropicalis NICM 3556 Fusarium
species Graphiumsp FIB4 Ochromonasdanica Aspergillus
awamori NRRL3112 |
|
Yeast |
Phanerochaete Rhodococus Rhodotorula Sphigmonas Trichosporon |
Phanerochaetechrysosporium Rhodococuserythropolis UPV-1 Rhodotorulacreatinivora Sphigmonaschlorophenica R4 2 Trichosporon species LE3 TrichosporonCutaneum R57 |
Table 2
|
Table 2 Reactors Used in the Phenol Degradation Khazi Et Al. (2010) |
|||
|
S.No |
Reactor |
Organisms Used |
Effect On Phenol
Degradation |
|
1 |
Packed bed Reactor |
Rhodococcus erythropolis |
Able to degrade completely phenol in defined mineral medium at a maximum
rate of 18kg of phenol m-3 per day |
|
2 |
Air stirred Reactor. |
Rhodococcus erythropolis UPV-1 |
Completely degrade phenol in synthetic wastewater at a volumetric productivity of 11.5 kg of phenol/m3 /day |
|
3 |
Packed bed Reactor |
Alcaligenes xylosoxidans
Y234 |
Able to degrade phenol of 1000 ppm completely in 60 h |
|
4 |
Hallow Fiber Membrane bioreactor |
Pseudomonas putida |
Able to degrade phenol of 1000 – 2000 mg/L |
|
5 |
Rotating biological contactors (RBC) |
Mixed culture |
Input loading 1754 – 3508 mg phenol/m2h |
|
6 |
Air lift bioreactor |
Alcaligenes xylosoxidans and Xanthomonas
maltophilia |
The fractional conversion of phenol over 99% was achieved |
|
7 |
Loop airlift bioreactor with a packed bed. |
Pseudomonas putida ATCC 17484 |
100% phenol removal was achieved at phenol loading rates up to 33120 mg /h m |
|
8 |
Pulsed plate bioreactor |
Immobilized Nocardia hydrocarbonoxydans |
100% degradation could be achieved with 300 and 500ppm influent phenol concentrations and at very low dilution. rate of 0.4094 1/h |
|
9 |
Self-cycling Fermentation in a stirred tank reactor |
Pseudomonas putida |
Substrate utilization rates as high as 14.5 kg of phenol per cubic meter
of fermentor volume per day of Fermentation, |
|
10 |
Granular activated carbon was incorporated into hollow fiber membrane bioreactor. |
Pseudomonas putida |
1000 ppm phenol was removed within 25 h. |
5. EFFECTS OF PARAMETERS FOR BIODEGRADATION OF PHENOL
5.1. Effect of pH
It is
believed that the internal environments of all living cells are fairly neutral. A pH of 4.0 or above is inhospitable to the majority of life. At pH 4.0 or 9.0, acids and bases
typically don't dissociate from one another and are resistant to electrostatic
fields, making them easier to enter cells. The optimal pH for phenol breakdown
in Pseudomonas putida NICM 2174 is 7.0. Annadurai et al. (2002).
5.2. Effect of temperature
The
key factor in the breakdown of organic contaminants is temperature, not the
presence of nutrients. Numerous investigations revealed that the rate of phenol
biodegradation considerably decreased around 30 °C Pakuła et al. (1999). The majority of studies on
phenol degradation have been done in the laboratory at the ideal temperature of
30°C, and they also found that as the temperature increased from 30 to 34°C, no
phenol degradation happened due to cell death, demonstrating that phenol
decomposition is a temperature-dependent process Annadurai et al. (1999). Growth rates typically double
for every 10°C increase in temperature in the typical mesophilic working range
of 10 to 30°C.The denaturation of proteins at higher temperatures lowers
mesophile growth rates, but growth rates in general do not vary between 35°C
and 40°C Ratkowsky et al. (1982).
5.3. EFFECT OF ADDITIONAL CARBON SOURCES ON PHENOL DEGRADATION
Diverse
strategies have been put out to handle highly concentrated phenolic wastewater
by discovering ways to get around substrate inhibition. These include the use
of genetically engineered microorganisms, cell immobilisation, and phenol
concentration adaption. One may be able to boost the cells' tolerance to
substrate inhibition by supplementing the growth medium with additional
conventional carbon sources, like yeast extract or glucose. Yeast extract has
also been seen to boost Pseudomonas putida's affinity
for phenol Armenante et al. (1995).
5.4. Effect of dissolved oxygen concentration
Most
frequently, oxygen serves as the ultimate electron acceptor in aerobic respiration
in aerobic microorganisms. Additionally, the microbial breakdown of a variety
of organic molecules, including hydrocarbons and compounds with aromatic rings,
requires a co-substratum known as molecular oxygen. The rate of organic load
breakdown under aerobic growth conditions is mostly determined by the dissolved
oxygen (DO) level. In studies on this subject, much attention has been paid to
how dissolved oxygen content affects microbial growth and respiration rate.
6. ADVANCES IN BIODEGRADATION OF PHENOL: KINETICS, MODELING, AND MASS TRANSFER
6.1. Kinetics and Modelling
Studies
on the kinetics of biodegradation reactions give an indication of how
effectively the microbial system is working. Gaining an understanding of these
dynamics will improve phenol removal efficiency and process control. By tying a
certain biomass growth rate to a specific substrate consumption rate, any
biodegradation process may be predicted (contaminant). A variety of kinetic
models have been used to describe the kinetics of microbial growth on phenol Table 3.
Two
of the most popular models for the biodegradation of phenol are the Monod model
Table 3, Equation 1 and the Haldane (Andrew's) model
Table 3, Equation 2.
Table 3
|
Table 3 Biodegradation Models (Kinetics and Mass Transfer) Taghreedalkhalid et al. (2012) |
|
|
Name
of model |
Equation |
|
Monad Haldane Linearized haldane Han- Levenspiel Yano Edwards Wang-Loh2 |
μ =
K is a constant
|
|
Monod: Sum Kinetics Binary mixture, no interaction Monod: sum kinetics Binary mixture, purely competitive interaction (inhibition) Binary mixture, Non-competitive inhibition Binary mixture, uncompetitive enzyme inhibition SKIPb Binary mixture unspecified type of interaction SKIP Three compound mixture, unspecified type of interaction Proposed by jiang et al. Michaelis-MentenC JD -factor Fick’s Lawd Thiele Moduluse |
|
Kp is a proportionality constant,
and aR is the specific substrate consumption rate
(mg/mg.hr) and ks is the saturation constant for
substrate consumption (mg/l), respectively. F(i)
depicts the functional relationship of the effect of metabolic intermediates on
phenol degradation. The interaction parameter, bIij,
shows how much substrate I influences substrate j's deterioration. v is the
reaction's initial speed (in mg/s), Km is the Michaelis constant, and V m is
the reaction's maximal speed in the case of catechol dioxygenase activity. ρp and De are the density of dried microbe (g/cm3) and
effective diffusion coefficient of phenol within the bead, respectively. dC is the phenol concentration within the immobilised
particles (mg/l), r is the radial position within the
bead. The real rate of deterioration is v (mg/g/hr). The rate constant, k', is
equal to kρp, where k is the fist-order
degradation rate constant (cm3/g.hr), and er0 is the particle's radius.
6.2. TEMPERATURE MODELS FOR KINETIC PARAMETERS
There
are fundamentally two types of treatment models for Monad maximum specific
growth rate (1/h). The dependent variable is written as rate0.5 in square-root models and as ln rate in Arrhenius type models. The
Arrhenius (1889) model's equation is
m = Ae - RT H* Equation 3
Where
m is the monad maximum specific growth rate (1/h) and H * is the Arrhenius
temperature characteristic (kJ/mol). It contains the generally assumed constant
temperature characteristic H*, however for microbial cultures, H* can vary by
up to three or four times, depending on the temperature range utilised.
6.3. Response Surface Methodology (RSM)
RSM,
is the most popular statistical method for optimising bioprocesses. It is a
graphical statistical method and empirical modelling methodology that is used
to assess the correlation between a number of
experimental variables that can be controlled and the observed findings.
Basically, this optimization approach entails three key steps: carrying out
statistically planned trials, determining the coefficies
in a mathematical model, and predicting the response and evaluating the model's
suitability. Prior understanding of the procedure to produce a statistical
model is necessary for response surface methodology, and takes the following
form:
Y = f (x1, x2, x3, x4,..., xk) Equation 4
The
variable xi is coded for statistical calculations as Xi = (xi -x0)/x.
The
following quadratic (second-degree) polynomial equation can be used to
approximate how these variables respond mathematically:
Y = b0 + bi Xi + bii Xi2 + bi j Xi Xj Equation 5
where
Y represents the anticipated response, b0 the offset term, bi the linear
effect, bii the square effect, and bij the interaction effect. The low, middle, and high
values of each variable (equally spaced) are denoted by the numbers -1, 0, and
+1, respectively. RSM was used by Annadurai et al. to optimise the medium
composition for P.putida's
phenol degradation (ATCC 31800).A mathematical model was then created to
illustrate the impact of each medium composition and their interactions on the
biodegradation of phenol. The mathematical expression of the relationship
between phenol degradation and variables like glucose, yeast extract, ammonium
sulphate, and sodium chloride was discovered. The
response for the aforementioned variables may always
be predicted by this model Taghreed al-khalid et al. (2012).
6.4. Mass Transfer
But the
inherent development It is inferred that models for the dynamics of cells in
suspension will also work well for biofilm cultures since kinetic models for
phenol degradation in biofilms are complex and difficult to create. Kinetics
degrade when the internal diffusion resistances are ignored. Despite its many
advantages, the fundamental drawback of biomass immobilisation by entrapment is
the limiting of product or nutrient diffusion caused by the resistance of the
protective framework. The effectiveness of deterioration is usually decreased
by a prevalent problem called diffusion limitation. The biomass is not easily
accessible to contaminants because the majority of
locations will be inside the bead. A support material should be strong,
chemically inert, and inexpensive to efficiently immobilise biomass. It should
have a strong connection to the cells, a large capacity for loading, and a
flexible structure to allow for minimal restriction on diffusion.
For
any phenol biodegradation process in a moving bed reactor using immobilised
biomass, there are three basic steps that occur in the bioreactor:
1) the movement of oxygen into the
liquid phase from the gas phase.
2) moving phenol, oxygen, and other
nutrients from the bulk liquid phase to the biofilm's surface; and
3) the biofilm's phenol, oxygen,
and other nutrients diffuse and react at the same time.
The
final step (3), which is a molecular phenomenon, is independent of the
reactor's flow parameters or turbulence.100 When it comes to the first process
(1), dissolved oxygen is a key factor. Oxygen transfer restrictions can cause
insufficient oxygen to have an impact on phenol biodegradation.
If
oxygen were utilised instead of air, there would be five times
increase in the oxygen mass transfer rate. Alternately, increasing stirring speed correspondingly
improved phenol decomposition and increased the mass transfer coefficient of
oxygen.
In
step (2), the substrate is thought to be transferred into the biofilm in two
steps:
·
the
substrate's transition from bulk liquid to a bioparticle's surface; and
·
Diffusion
through the layer of microorganisms (biofilm).
Numerous
research have concentrated on this process since the
speed at which phenol is transported from the bulk phase to the biofilm's
surface would directly affect the biochemical reaction taking place there. The
design and modelling of bioreactors must therefore take into account external
mass transfer coefficients for the transport of phenol from the bulk phase to
the surface of the biofilm.
7. Conclusion
Phenolic compounds must be eliminated as priority pollutants in order to maintain environmental quality. Since biological treatment is the most successful, cost-effective, and ecologically friendly technology now available, it is receiving more and more attention in the field of pollution control. The basic function of microbial metabolism is energy conversion, and it is controlled by enzymatic systems in which reaction intermediates are important. In addition, numerous models have been put forth in order to better understand the kinetics of phenols' biodegradation. The Haldane and Monod models are most frequently employed. The effectiveness of the biodegradation process is significantly influenced by the mass transport mechanisms and regimes.
8. Future scope and directions
Development
efforts should focus on novel types of bioreactors targeting practical
utilization and efficient long-term performance. It is essential to develop and design efficient reactors that
would reduce mass transfer limitations and enhance the degradation rate. Novel
processes are required as well.
Credit author statement
M.N.V.
Sai Rachana: Organic pollution of phenol, sources of phenol, Limits of phenol; J.
Akhila : harmful effects of phenol ; L. Reshma: analysis of phenol, degradation of
phenol through aerobic and anaerobic pathways M. Yamini : Mechanism of phenol, Effects of Parameters
for biodegradation of phenol, metabolic pathways for phenol : Supervision.
V.
Sridevi: Advances in biodegradation of
phenol: kinetics, modelling and Mass Transfer, Various microorganisms, and
reactors for phenol biodegradation , visulisation : Husam Talib Hamzah
, Grammar and spellcheck – R.Sri Harsha and Katru Ramya sugandhi.
CONFLICT OF INTERESTS
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
ACKNOWLEDGMENTS
The authors are sincerely thankful to the Department of Chemical Engineering, Andhra University, Visakhapatnam, Andhra Pradesh, India.
REFERENCES
Agarry,
S. E., Durojaiye, A. O., and Solomon, B. O. (2008). Microbial
Degradation of Phenols : A Review. International Journal of Environment and
Pollution, 32(1), 12–28. Https://Doi.Org/10.1504/Ijep.2008.016895.
Aisami, A., Yasid, N. A., and Shukor, M. Y. A. (2020). Optimization of Cultural and Physical Parameters for Phenol Biodegradation by Newly Identified Pseudomonas Sp. Aq5-04. Journal of Tropical Life Science, 10(3), 223–233. Https://Doi.Org/10.11594/Jtls.10.03.06
Al-Khalid, T., and El-Naas, M. H. (2012). Aerobic
Biodegradation of Phenols : A Comprehensive Review. Critical Reviews In
Environmental Science and Technology, 42(16), 1631–1690.
Https://Doi.Org/10.1080/10643389.2011.569872.
Alqahtani,
A., Tongkao-On, W., Li, K. M., Razmovski-Naumovski, V., Chan, K., and Li, G. Q.
(2015). Seasonal Variation of Triterpenes and Phenolic Compounds In
Australian Centella Asiatica (L.) Urb. Phytochemical Analysis, 26(6), 436–443.
Https://Doi.Org/10.1002/Pca.2578.
Annadurai, G., Juang, R. S., and Lee, D. J. (2002).
Microbiological Degradation of Phenol Using Mixed Liquors of Pseudomonas Putida
and Activated Sludge. Waste Management, 22(7), 703–710.
Https://Doi.Org/10.1016/S0956-053x(02)00050-8.
Annadurai,
G., Mathalai Balan, S., and Murugesan, T. (1999). Box-Behnken Design In
The Development of Optimized Complex Medium For Phenol Degradation Using
Pseudomonas Putida (Nicm 2174). Bioprocess Engineering, 21(5), 415–421.
Https://Doi.Org/10.1007/Pl00009082.
Armenante,
P. M., Fava, F., and Kafkewitz, D. (1995). Effect Of Yeast Extract on
Growth Kinetics During Aerobic Biodegradation of Chlorobenzoic Acids.
Biotechnology and Bioengineering, 47(2), 227–233.
Https://Doi.Org/10.1002/Bit.260470214.
Bhatia, S. K., Joo, H. S., and Yang, Y. H. (2018). Biowaste-To-Bioenergy Using Biological Methods – A Mini-Review. Energy Conversion And Management, 177, 640–660. Https://Doi.Org/10.1016/J.Enconman.2018.09.090.
Coskun,
O. (2016). Separation Techniques : Chromatography. Northern Clinics of
Istanbul, 3(2), 156–160. Https://Doi.Org/10.14744/Nci.2016.32757.
Da
Silva Siqueira, E. M., Félix-Silva, J., De Araújo, L. M., Fernandes, J. M.,
Cabral, B., Gomes, J. A., De Araújo Roque, A., Tomaz, J. C., Lopes, N. P., De
Freitas Fernandes-Pedrosa, M., Giordani, R. B., and Zucolotto, S. M. (2016).
Spondias Tuberosa (Anacardiaceae) Leaves : Profiling Phenolic Compounds By
Hplc-Dad and Lc-Ms/Ms and In Vivo Anti-Inflammatory Activity. Biomedical
Chromatography, 30(10), 1656–1665. Https://Doi.Org/10.1002/Bmc.3738.
Duan, W., Meng, F., Cui, H., Lin, Y., Wang, G., and Wu, J. (2018).
Ecotoxicity of Phenol and Cresols to Aquatic Organisms : A Review.
Ecotoxicology and Environmental Safety, 157, 441–456. Https://Doi.Org/10.1016/J.Ecoenv.2018.03.089.
Evans,
W. C. (1947). Oxidation of Phenol and Benzoic Acid by Some Soil
Bacteria. Biochemical Journal, 41(3), 373–382. Https://Doi.Org/10.1042/Bj0410373.
Fan, J., Fan, Y., Pei, Y., Wu, K., Wang, J., and Fan, M. (2008). Solvent Extraction of Selected Endocrine-Disrupting Phenols Using Ionic Liquids. Separation and Purification Technology, 61(3), 324–331. Https://Doi.Org/10.1016/J.Seppur.2007.11.005.
González, G., Herrera, G., García, M. T., and Peña, M. (2001). Biodegradation of Phenolic Industrial Wastewater in a Fluidized Bed Bioreactor With Immobilized Cells of Pseudomonas Putida. Bioresource Technology, 80(2), 137–142. Https://Doi.Org/10.1016/S0960-8524(01)00076-1.
Harwood, C. S., and Parales, R. E. (1996). The
Beta-Ketoadipate Pathway and the Biology of Self-Identity. Annual Review of
Microbiology, 50, 553–590. Https://Doi.Org/10.1146/Annurev.Micro.50.1.553.
Kazemi, P., Peydayesh, M., Bandegi, A., Mohammadi, T., and
Bakhtiari, O. (2014). Stability and Extraction Study of Phenolic
Wastewater Treatment by Supported Liquid Membrane Using Tributyl Phosphate and
Sesame Oil as Liquid Membrane. Chemical Engineering Research and Design, 92(2),
375–383. Https://Doi.Org/10.1016/J.Cherd.2013.07.023.
Kilby, B. A. (1948). The Bacterial Oxidation of Phenol to Beta-Ketoadipic Acid. Biochemical Journal, 43(1), V–V.
Kuc, M. E., Azerrad, S., Menashe, O., and Kurzbaum, E. (2022).
Efficient Biodegradation of Phenol At High Concentrations By Acinetobacter
Biofilm At Extremely Short Hydraulic Retention Times. Journal Of Water Process
Engineering, 47, 102781. Https://Doi.Org/10.1016/J.Jwpe.2022.102781.
Letelier, M. E., Rodríguez-Rojas, C., Sánchez-Jofré, S., and Aracena-Parks, P. (2011). Surfactant and Antioxidant Properties of an Extract From Chenopodium Quinoa Willd Seed Coats. Journal of Cereal Science, 53(2), 239–243. Https://Doi.Org/10.1016/J.Jcs.2010.12.006.
Mocan, A., Crișan, G., Vlase, L., Crișan, O., Vodnar, D.
C., Raita, O., Gheldiu, A. M., Toiu, A., Oprean, R., and Tilea, I. (2014).
Comparative Studies on Polyphenolic Composition, Antioxidant and Antimicrobial
Activities of Schisandra Chinensis Leaves and Fruits. Molecules, 19(9),
15162–15179. Https://Doi.Org/10.3390/Molecules190915162.
Naczk, M., and Shahidi, F. (2006). Phenolics In Cereals,
Fruits and Vegetables: Occurrence, Extraction and Analysis. Journal of
Pharmaceutical and Biomedical Analysis, 41(5), 1523–1542.
Https://Doi.Org/10.1016/J.Jpba.2006.04.002.
Pakuła, A., Bieszkiewicz. E., Boszczyk-Maleszak, H., and Mycielski, R. (1999). Biodegradation of Phenol by Bacterial Strains from Petroleum-Refining Wastewater Purification Plant. Europepmc.Org.
Patil, S. S. (2014). Biodegradation Study of Phenol By Burkholderia Sp. Ps3 And Bacillus Pumilus Os1 Isolated From Contaminated Soil.
Pouraboli, I., Nazari, S., Sabet, N., Sharififar, F., and
Jafari, M. (2016). Antidiabetic, Antioxidant, and Antilipid Peroxidative
Activities Of Dracocephalum Polychaetum Shoot Extract In Streptozotocin-Induced
Diabetic Rats: In Vivo and in Vitro Studies. Pharmaceutical Biology, 54(2),
272–278. Https://Doi.Org/10.3109/13880209.2015.1033561.
Ratkowsky, D. A., Olley, J., Mcmeekin, T. A., and Ball, A.
(1982). Relationship Between Temperature and Growth Rate of Bacterial
Cultures. Journal of Bacteriology, 149(1), 1–5.
Https://Doi.Org/10.1128/Jb.149.1.1-5.1982.
Rejczak, T., and Tuzimski, T. (2017). Application of
High-Performance Liquid Chromatography With Diode Array Detector for
Simultaneous Determination of 11 Synthetic Dyes In Selected Beverages and
Foodstuffs. Food Analytical Methods, 10(11), 3572–3588.
Https://Doi.Org/10.1007/S12161-017-0905-3.
Rozich,
A. F., and Colvin, R. J. (1986). Effects of Glucose on Phenol
Biodegradation by Heterogeneous Populations. Biotechnology and Bioengineering,
28(7), 965–971. Https://Doi.Org/10.1002/Bit.260280706.
Sankhalkar,
S., and Vernekar, V. (2016). Quantitative and Qualitative Analysis of
Phenolic and Flavonoid Content In Moringa Oleifera Lam and Ocimum Tenuiflorum
L. Pharmacognosy Research, 8(1), 16–21.
Https://Doi.Org/10.4103/0974-8490.171095.
Satish,
K., Neeraj, Viraj, K. M., and Santosh, K. K. (2018). Biodegradation of
Phenol By Free and Immobilized Candida Tropicalis Npd1401. African Journal of
Biotechnology, 17(3), 57–64. Https://Doi.Org/10.5897/Ajb2017.15906.
Sun, J., Mu, Q., Kimura, H., Murugadoss, V., He, M., Du, W., and Hou,
C. (2022). Oxidative Degradation of Phenols and Substituted Phenols In
The Water And Atmosphere : A Review. Advanced Composites and Hybrid Materials,
5(2), 627–640. Https://Doi.Org/10.1007/S42114-022-00435-0.
Touliabah, H. E. S., El-Sheekh, M. M., Ismail, M. M., and
El-Kassas, H. (2022). A Review of Microalgae- and Cyanobacteria-Based
Biodegradation of Organic Pollutants. Molecules, 27(3).
Https://Doi.Org/10.3390/Molecules27031141.
Vaičiulyte, V., Butkienė, R., and Ložiene, K. (2016). Effects of Meteorological Conditions and Plant Growth Stage On the Accumulation of Carvacrol and its Precursors in Thymus Pulegioides. Phytochemistry, 128, 20–26. Https://Doi.Org/10.1016/J.Phytochem.2016.03.018.
Williams,
R. J., and Evans, W. C. (1975). The Metabolism of Benzoate By Moraxella
Species Through Anaerobic Nitrate Respiration. Evidence For a Reductive
Pathway. Biochemical Journal, 148(1), 1–10. Https://Doi.Org/10.1042/Bj1480001a.
Xiang, Y., Xing, Z., Liu, J., Qin, W., and Huang, X. (2020). Recent Advances In The Biodegradation of Polychlorinated Biphenyls. World Journal of Microbiology and Biotechnology, 36(10), 145. Https://Doi.Org/10.1007/S11274-020-02922-2.
 This work is licensed under a: Creative Commons Attribution 4.0 International License
This work is licensed under a: Creative Commons Attribution 4.0 International License
© Granthaalayah 2014-2023. All Rights Reserved.